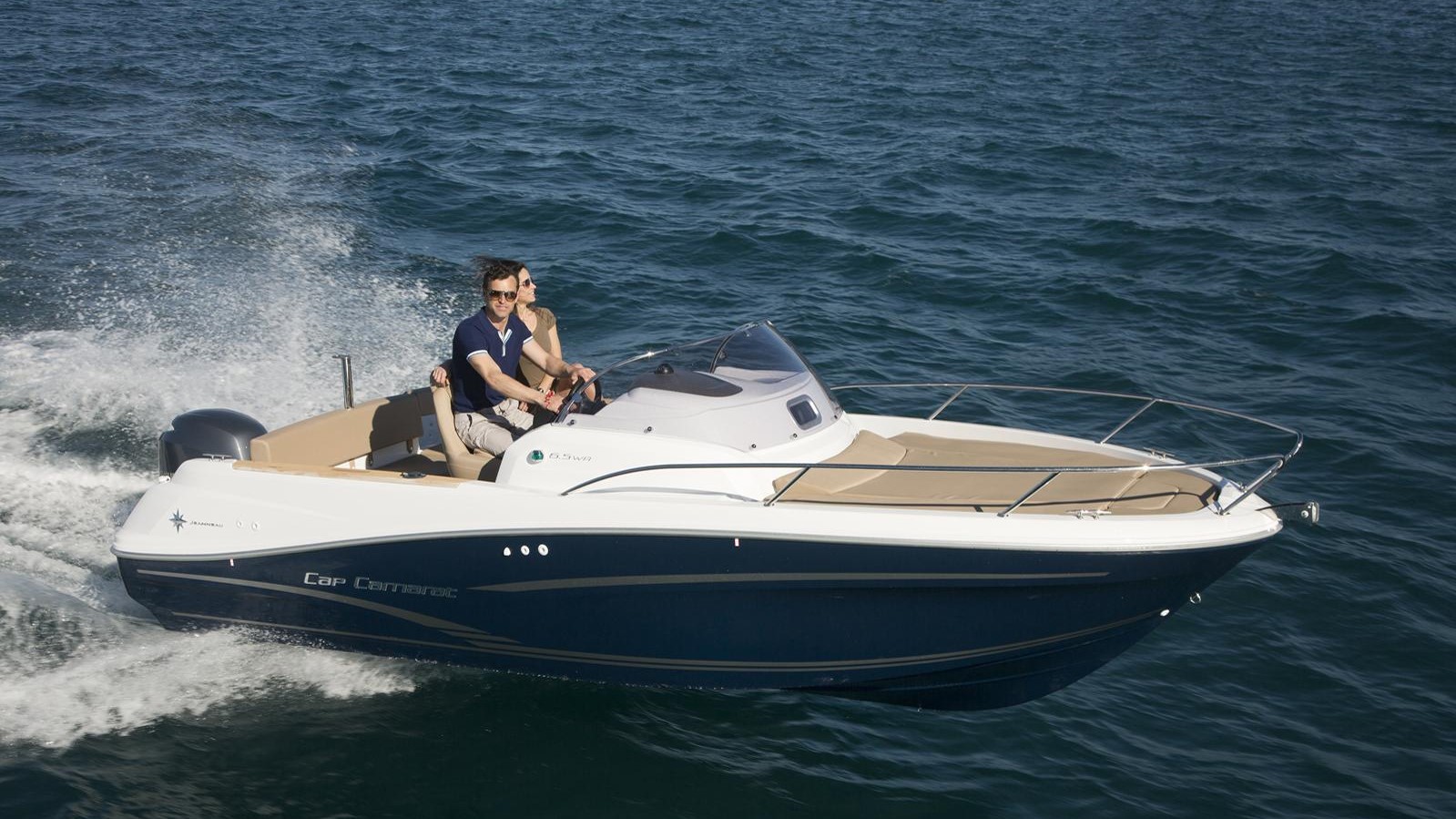

We implemented a functionality to compute assignment with two reference databases. Taxonomic assignments of ASVs are carried out by IDTAXA (part of the DECIPHER package), an algorithm based on a machine learning method 5. This step results in an object with representing sequences, a raw ASV counts table and a text file containing statistics from the denoising process. ASV identifiers are sequences translated into MD5 hashes which are unique identifiers based on the DNA sequences offering the possibility to be compared between projects. For 16S amplicon, primers are trimmed based on the primer sequence length.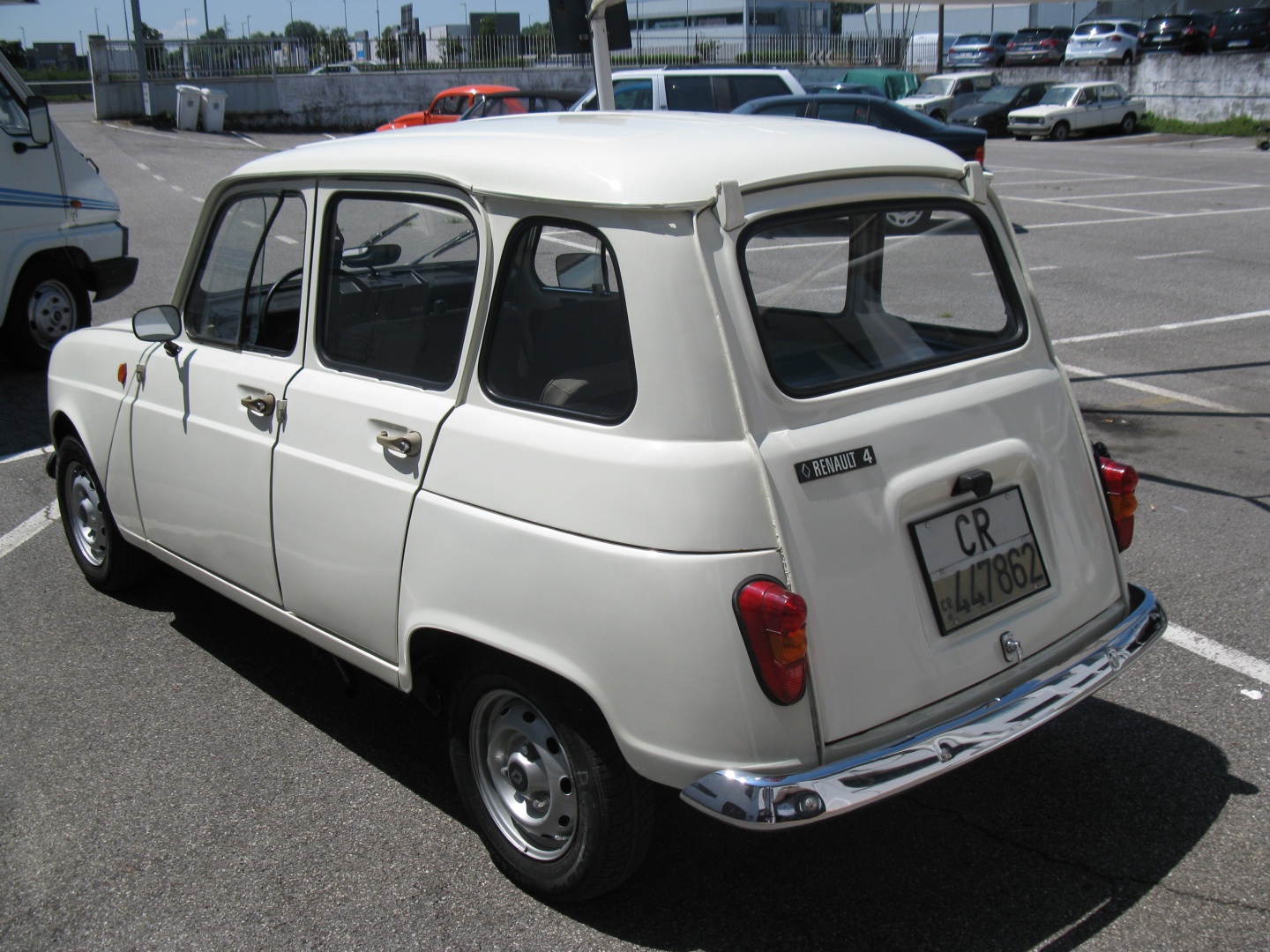 
rANOMALY handles processing any region of 16S (V1 to V9) and ITS amplicon (ITS1, ITS2) sequences, in which an additional step of cutadapt 4 removes ITS probes left in some short sequences. This improves resolution of the potential presence of microbial organisms by using a prediction model to correct sequencing errors before aggregating similar sequences. The denoising process is handled using the dada2 R package 3 which produces amplicon sequence variants (ASV) as a taxonomic unit. Samples must have been previously demultiplexed into one file per sample with the file name following this syntax: _R.fastq. It can produce high quality figures for Rmarkdown reports along with statistical tests ready for publication. 
rANOMALY only needs a CSV table describing the metadata for each sample, and a folder containing the corresponding fastq files as input. The package allows the workflow to be executed on any R environment. rANOMALY is fully implemented in R language in which each step correspond to one function, allowing to easy implementation of new features or tools while being easy to use and maintain. Here we present rANOMALY, a scalable and lightweight R package which is able to handle every step of a metabarcoding analysis, from read cleaning, contaminant filtering, taxonomic assignment, to advanced statistical analysis. Methods and software are continuously evolving and the main challenge for bioinformaticians is to implement the most recent and effective ones in their analysis. Metabarcoding generates a large amount of data and a lot of applications already exist for their processing (FROGS 1, qiime 2). Studies of microbial communities tends to become a daily routine analysis for lots of laboratories and the main method to explore microbial diversity is metabarcoding, which is an amplicon targeted sequencing method (16S for bacteria and ITS for fungi). Our package produces ready to publish figures, and all of its outputs are made to be integrated in Rmarkdown code to produce automated reports. It integrates all assets of the latest bioinformatics methods, such as better sequence tracking, decontamination from control samples, use of multiple reference databases for taxonomic annotation, all main ecological analysis for which we propose advanced statistical tests, and a cross-validated differential analysis by four different methods. rANOMALY is an easy to install and customizable R package, that uses amplicon sequence variants (ASV) level for microbial community characterization. The main purpose of this package is to automate bioinformatic analysis, ensure reproducibility between projects, and to be flexible enough to quickly integrate new bioinformatic tools or statistical methods. This workflow is based on several R functions and performs automatic treatments from fastq sequence files to diversity and differential analysis with statistical validation. We present an R package for data analysis of 16S and ITS amplicons based sequencing. Bioinformatic tools for marker gene sequencing data analysis are continuously and rapidly evolving, thus integrating most recent techniques and tools is challenging.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
